Yahya Chaudry
Harvard Staff Writer

Xiaowei Zhuang’s lab made the invisible visible — and changed how scientists study health and disease
Harvard Staff Writer
Throughout the history of biology, numerous discoveries stemmed from observing living systems, but seeing the innerworkings of living organisms presents challenges.
Xiaowei Zhuang, David B. Arnold Professor of Science, has spent her career developing imaging technologies that enable visualization of minuscule objects in living systems. With a toolbelt of powerful genomic and imaging tools, she has made it possible to see, count, and map molecules inside cells and tissues with unprecedented clarity and scale — reshaping how scientists can study everything from synapses to entire brains.
“I love this quote: ‘Progress in science depends on new techniques, new discoveries, and new ideas. Probably in that order’,” said Zhuang, who frames her philosophy from Nobel laureate Sydney Brenner. “It signifies the importance of technology development. New technologies can open entirely new venues, enable previously intractable studies, and drastically accelerate discoveries.”
Zhuang’s research sits at the intersection of physics, chemistry, and biology, seeking to expand the frontiers of each field. She trained first and foremost as a physicist. After earning her bachelor’s degree in physics from the University of Science and Technology of China in 1991, she moved to the United States for graduate school at the University of California, Berkeley, where she studied nonlinear optics with Yuen‑Ron Shen. From there, she joined the lab of Nobel laureate Steven Chu at Stanford as a postdoctoral fellow, using imaging-based tools to watch single molecules in action.

When she arrived at Harvard in 2001, Zhuang started a research program developing advanced imaging methods to study biomolecules and molecular assemblies in their native context, living cells. For example, she developed single-particle tracking methods to study how viruses infect cells. Although these studies generated interesting insights, a major limitation in light microscopy — the diffraction limit — significantly hindered the research. Because of the diffraction of light waves, the resolution of conventional microscopes was limited to about half the wavelength of light, obscuring the details of molecular interactions in cells and tissues.
By 2006, Zhuang and her colleagues had devised a way to overcome this barrier. The approach, called stochastic optical reconstruction microscopy (STORM), uses light‑switchable fluorescent dyes so that only a sparse subset of labeled molecules glows at any given time, allowing their positions to be precisely recognized. By repeatedly turning different subsets on and off and localizing each molecule with nanometer precision, STORM reconstructs super‑resolved images far beyond the diffraction limit.
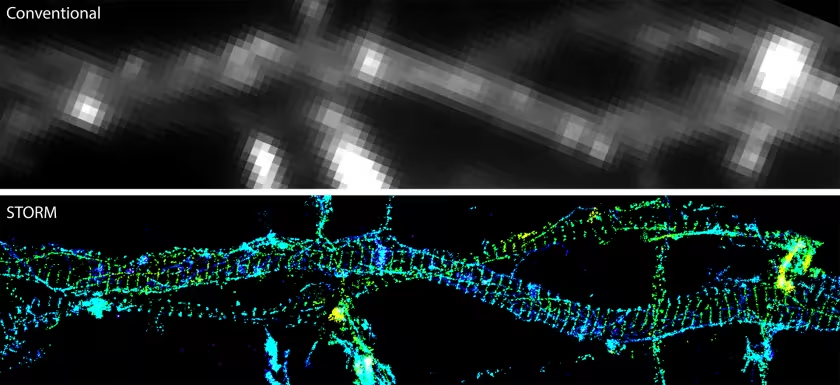
At first, STORM was a way to see known structures in finer detail, but it soon became a way to discover entirely new structures in cells. Applying 3D STORM to neurons, Zhuang’s lab revealed a previously unknown, exquisitely regular lattice of actin, spectrin, and associated proteins along neurites — a periodic membrane skeleton structure that anchors ion channels and signaling molecules and reshaped thinking about axon and dendrite organization.
“It’s a really remarkable feeling,” Zhuang said, “a discovery of something that people never knew existed before.”
As STORM became widely used, Zhuang’s “restless” spirit prompted her to develop something entirely new again — this time taking imaging to the genome scale. Existing imaging methods could measure a few genes at a time in individual cells. Zhuang wanted to scale up to thousands. It was a task that appeared impossible because thousands of distinguishable colors in light microscopy don’t exist. “When STORM got mature, I got bored and wanted to do something entirely new,” she said. “How to take imaging to the genome scale became a problem I was obsessed with every night, until one night, the idea clicked — in a Eureka moment.”
The lightbulb was multiplexed error‑robust fluorescence in situ hybridization (MERFISH), which uses combinatorial imaging to solve the scaling problem, allowing the number of genes imaged to increase exponentially with the number of imaging rounds. Specifically, MERFISH assigns unique barcodes (i.e. 101001…1, 110001…1, etc.) to individual genes, reads those codes bit by bit over many rounds of imaging, and uses coding schemes capable of error detection and correction to ensure accurate identification. The method allows researchers to count and localize RNAs from thousands of genes in single cells in situ, producing genome‑scale, spatially resolved expression maps, as Zhuang reported in a 2015 Science paper.
“By studying one gene at a time, you could get the pieces” Zhuang said, “but you couldn’t get the whole picture. For that, we need to be able to image at the genome scale.”

With MERFISH, those whole‑picture views have become reality. It enables gene expression profiling of individual cells in intact tissues, which in turn allows systematic identification, spatial mapping, and functional interrogation of different types of cells in complex tissue. In a co-authored study with Catherine Dulac reported in Science in 2018, Zhuang used this approach to not only identify and map molecularly defined cell types in the mouse hypothalamic preoptic region, but also determine their functional roles in social behaviors, such as parent, mating and aggression, linking cell identity, spatial organization, and behaviorally relevant function.
Most recently, in a tour-de-force 2023 Nature paper, the Zhuang lab used MERFISH to create a molecularly defined, spatially resolved cell atlas of the entire adult mouse brain — imaging more than 1,100 genes in roughly 10 million cells and integrating those data with large‑scale single‑cell RNA‑sequencing. The result is a comprehensive map of over 5,000 transcriptionally defined cell populations and where they reside in the brain.
Such work has helped define the field of spatial omics and underpins global efforts like the Human Cell Atlas. Zhuang has also extended MERFISH to enable spatial multiomics, measuring not only the transcriptome, but also the 3D organization of the genome and the epigenome to understand regulatory mechanisms underlying gene expression. She also developed a MERFISH-based functional genomics method that allows pooled genetic screening for rich cellular phenotypes, accelerating discoveries of molecular mechanisms underlying cellular and tissue functions.
For Zhuang, who has been recognized with many of science’s top honors, including the Breakthrough Prize in Life Sciences, the National Academy of Sciences Award for Scientific Discovery, and the Lurie Prize in Biomedical Sciences, the point of developing new technology is not just to push technological boundaries, but to open new biological questions.
“Because there are major gaps in technology, there are a lot of problems that you just simply cannot study,” Zhuang said. “We are keen on technology development to allow us to see things that were just plainly not possible before.”
Zhuang and her lab members marvel at the interplay between technology development and scientific discoveries. In recent work in collaboration with MIT Professor of Biology Jonathan Weissman, reported in Cell, Zhuang used the imaging‑based pooled CRISPR screening platform to combine genetic perturbations with spatially resolved transcriptomic and imaging readouts, offering a powerful way to determine gene functions and link them to cell identity and tissue architecture at scale. Using this approach, the researchers identified novel genetic regulators of liver function and dysfunction.
“Collaborators like Catherine Dulac and Jonathan Weissman added a lot of pleasure to scientific research. We have collaborated on multiple different projects over the years,” Zhuang said. “I also feel very fortunate to have the privilege of supervising multiple generations of talented and dedicated lab members. Working with them is the most fun and rewarding part of doing science.”
With each new technique, from STORM’s super‑resolved images to MERFISH’s brain‑wide cell atlases and next‑generation perturbation platforms, Zhuang continues to show that when you change what you can see, you change what you can ask — and what you can discover.

Cap on A grades and shift from GPA to ‘average percentile rank’ set to go into effect fall 2027

A new AI tool called Empirical Research Assistance can automatically write high-performance scientific software, even outperforming software written by experts.

Hawaiʻi’s Alyssia Wiesenbauer arrived with an interest in studying volcanoes. She leaves with a deep knowledge of nature and museums.