Yahya Chaudry
Harvard Staff Writer
Interdisciplinary research on mice maps olfactory system
Harvard Staff Writer
Mice rely on odors to find mates, avoid predators, identify food sources, and recognize their own pups. Yet how those crucial scents are organized — and understood by the brain — has remained mysterious.
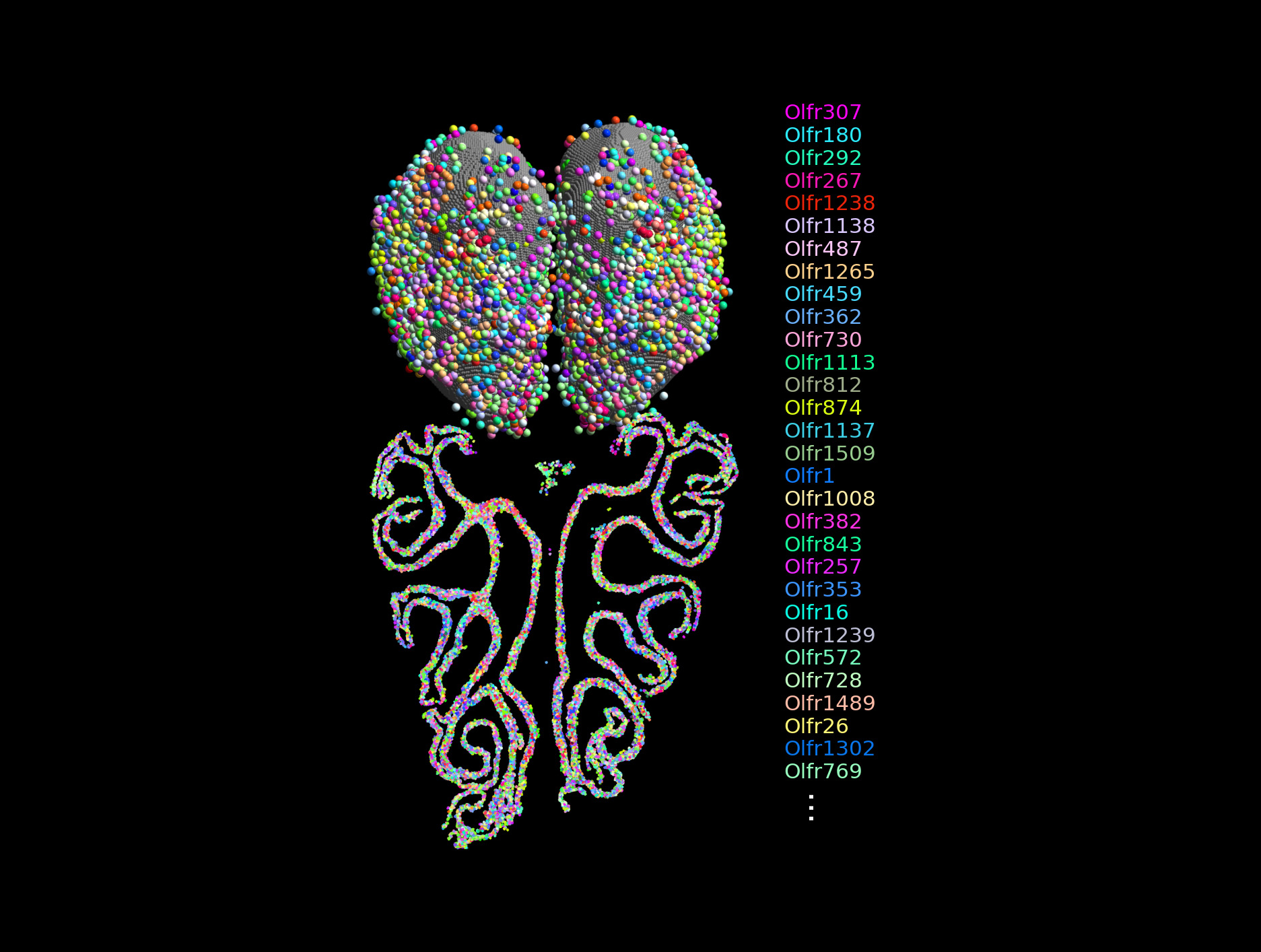
Now, a team in two labs has created the most comprehensive map to date of how the primary olfactory system detects and organizes social and predator odors. The work, published in Cell, shows that the mouse’s main smell pathway is arranged in smooth gradients that are mirrored between the nose and the olfactory bulb, and that cues from mates, pups, and predators activate distinct regions within that map.
The study was led jointly by Catherine Dulac, Xander University Professor in the Department of Molecular and Cellular Biology, and Xiaowei Zhuang, David B. Arnold Jr. Professor of Science in the Department of Chemistry and Chemical Biology. Both are Howard Hughes Medical Institute investigators.
Dulac, who has spent decades studying the neural basis of social behavior, likes to frame the puzzle with a question she credits to her former mentor, Nobel laureate Richard Axel: “How does the brain know what the nose knows?”
“You detect something in the nose, and somewhere the brain has to know what was detected in the olfactory epithelium,” she said. “There has to be a mode of organization such that your brain understands what’s happening in your nose. That organization has to be similar between you and me, such that we have similar percepts.”
The main olfactory epithelium expresses more than a thousand odorant receptor genes, many of them similar to each other. Traditionally, researchers could study only a few at a time.
“What you could do before,” Dulac said, “was take a few receptors and try to extrapolate the organization from those.”
But that left open the possibility that important patterns were being missed.
The breakthrough came from a technology pioneered in Zhuang’s lab called MERFISH — Multiplexed Error‑Robust Fluorescent In Situ Hybridization) a genome-scale imaging method that reads out expression of thousands of genes directly into intact tissue, preserving where each cell sits.
“In this work, we had to image about a thousand genes and then see where they were expressed in the olfactory system at the single‑cell level,” Zhuang said. “If you were to do one gene at a time and study a few genes, you really don’t get the whole picture. And MERFISH is a perfect technology for getting the holistic picture.”
Using specialized MERFISH probes, Bogdan Bintu, Ph.D. ’21, a former student in the Kenneth C. Griffin Graduate School of Arts and Sciences in both labs, produced with colleagues a near‑complete atlas of receptor expression in the nose and its projections into the olfactory bulb. Sensory neurons were found to be arranged in two layers in the nose, and their outgoing connections created the same layered pattern in the brain.
“By doing MERFISH on the olfactory bulb, we could map all the projections,” Dulac said. “Now we could have access to the map of the entire olfactory bulb with all these receptors. That was absolutely magnificent.
“There was a really beautiful match between the two organizations,” she added. “So now, we have a deep understanding of the organizational mode of the olfactory epithelium and olfactory bulb.”
A separate study led by Sandeep Robert Datta, professor of neurobiology in the Blavatnik Institute at Harvard Medical School, published consistent findings in the same issue of Cell.
To connect structure to function, the team exposed mice to naturalistic cues — cat bedding as a predator odor, male and female intruders as social cues, and litters of pups — and used MERFISH together with markers of neural activity to see which receptor‑defined neurons were engaged and where they projected. Moreover, the group also tested whether the sex and physiological states of mice affect their detection of scents. Remarkably, striking differences were uncovered in the detection of infants in lactating mothers compared to virgin females; several odorant receptions and separate regions of the olfactory bulb were activated when mothers smelled pups, but those same receptions stayed dormant in virgin females.
“Usually once you find these cells, where they are, and how they interact, you also care about their functional role,” Zhuang said. “This work already puts a functional role into this spatial atlas of cells, and that is, Which neurons are responsible for smelling which odor?”
“This is a powerful approach to study the function of cells at a systems level,” she added. The same combination of spatial mapping and activity readouts can be applied to other brain regions and behaviors, tracing how specific molecularly defined cells participate in neural circuits controlling other behaviors.
In the mice, Dulac noted, the distinctions uncovered in this atlas are existential.
“Mice function and survive in the world thanks to their olfactory system,” she said. “They need to detect who is a potential mate, what is a danger, what is a predator. Parents also need to identify infants with great accuracy and sensitivity in order to ensure their survival.”
From her perspective, the implications go well beyond smell.
“From a neuroscience standpoint, the fact that it’s olfaction in mice or vision in humans or auditory in a bird doesn’t really matter,” Dulac said. “Because essentially what you want to understand is the logic of how you recognize an object or an individual in the world, and how this logic is modulated according to your physiological needs. This is the holy grail of neuroscience.”
This research was supported by the National Institute of Mental Health.
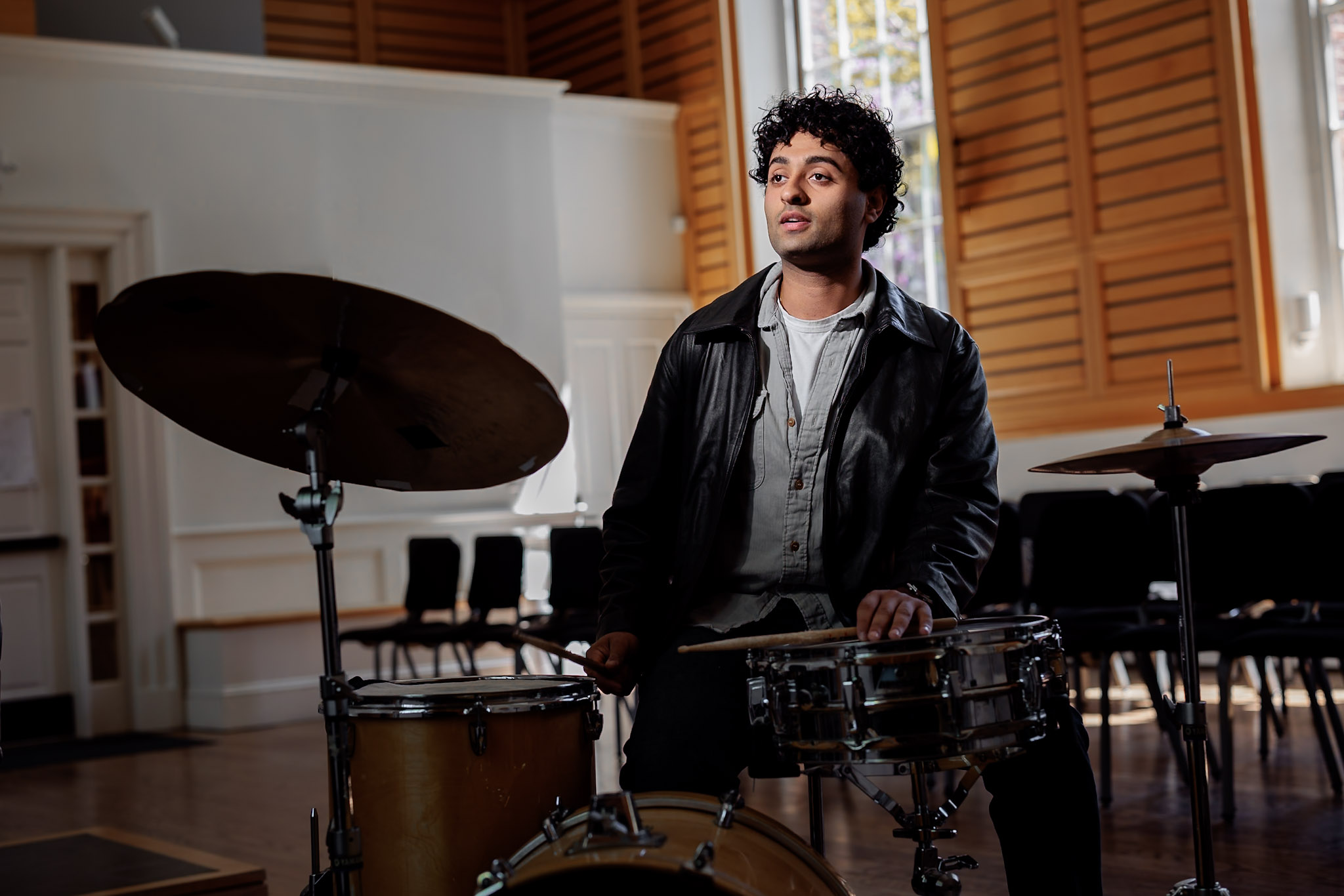

At Harvard, drummer Raghav Mehrotra ’26 built knowledge about the music industry — and a sizable social media following.
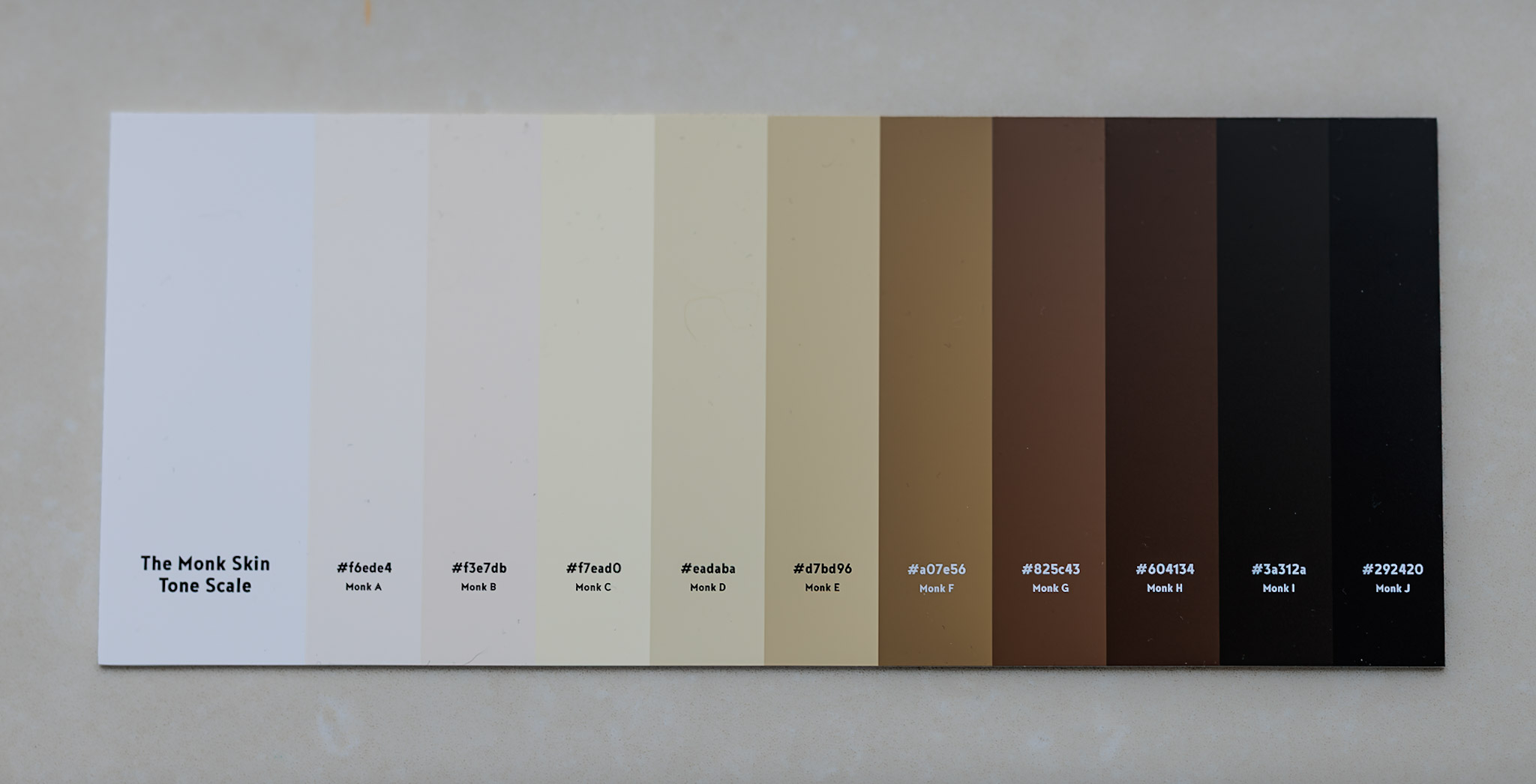

The Monk Skin Tone scale, devised in 2019 by Professor Ellis Monk, helps create more accurate medical diagnostic tools for patients with dark skin
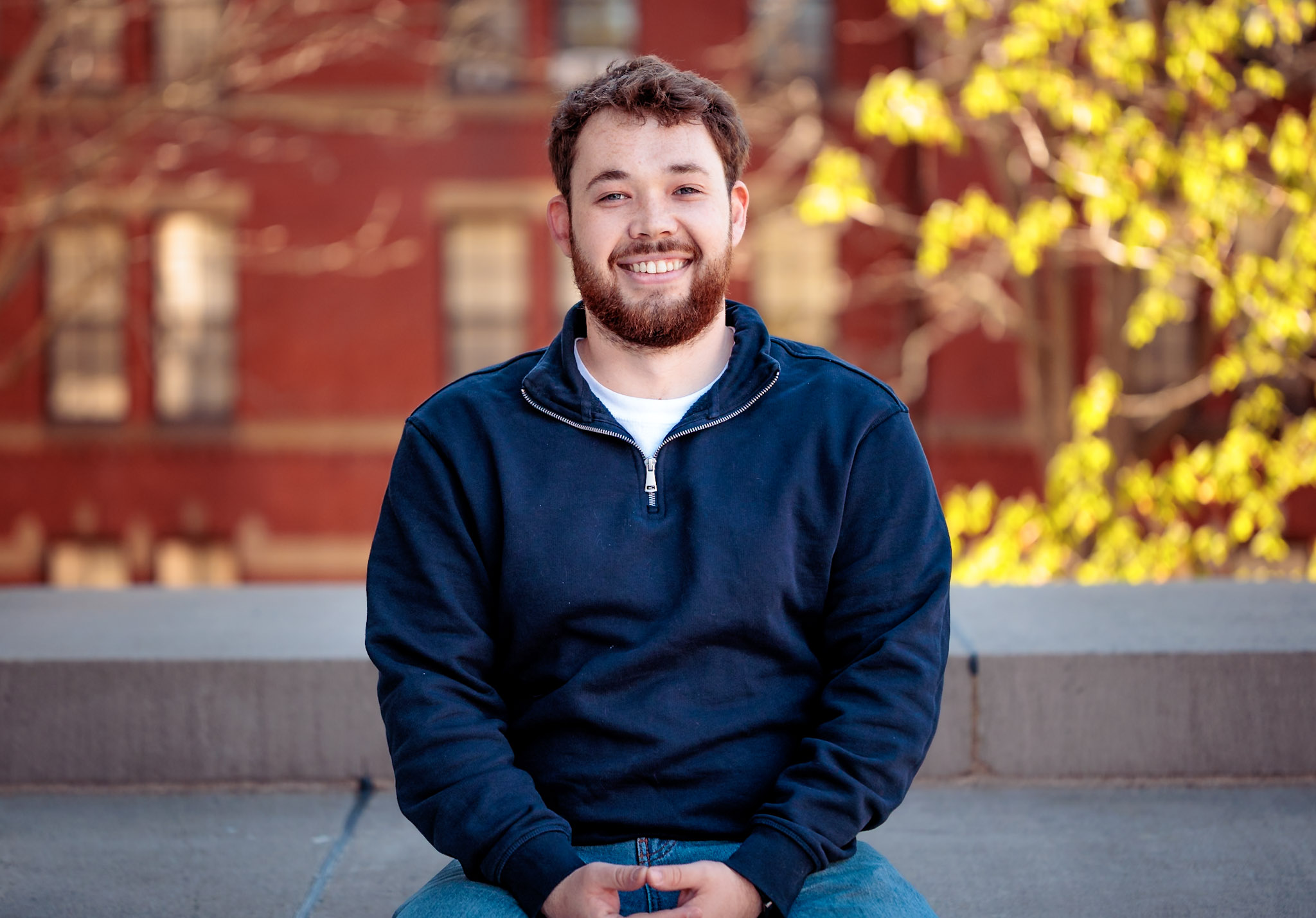

A few months shy of 30, Richard Glazunov is poised to graduate from Harvard College with a degree in government.